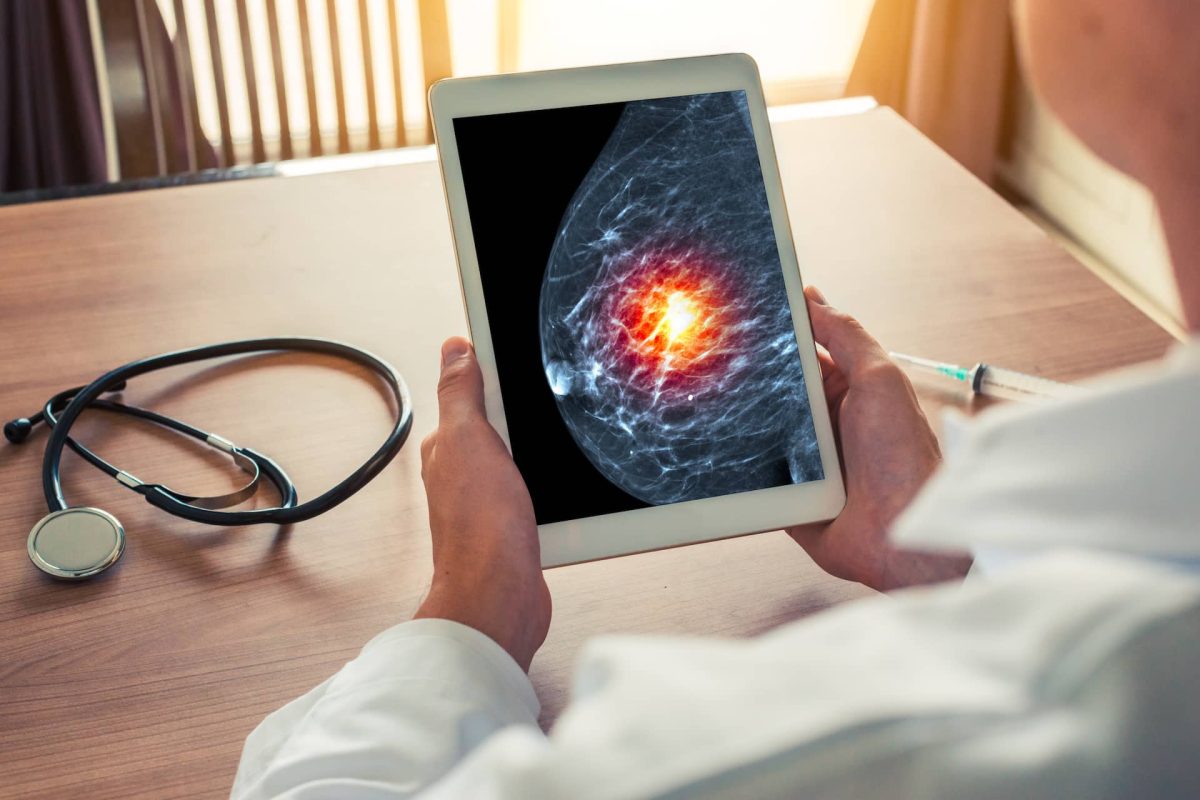
While routine screening has been shown to decrease breast cancer mortality in multiple randomized controlled trials, mammography is limited by subjective human interpretation. Recent advances in improved computer processing power, cloud-based data storage capabilities, and availability of large imaging datasets have led to renewed excitement for applying artificial intelligence (AI) to mammography interpretation. A collaboration between the University of Washington (Christoph Lee) and UCLA (Joann Elmore, Arash Naeim, William Hsu), this project is undertaking a unique academic-industry partnership to validate, refine, scale, and clinically translate our proven 2D mammography AI algorithm to 3D mammography interpretation. We are validating an existing algorithm for 2D mammography using UCLA’s Athena Breast Health Network. We are enhancing this 2D AI algorithm with expert radiologist supervision and will be examining the impact of adding novel non-imaging data parameters, including genetic mutation and tumor molecular subtype data, to help train the AI model to identify more clinically significant cancers.
We are using several novel technical algorithmic approaches to scale from 2D to 3D mammography that in our preliminary studies have shown improved accuracy beyond radiologist interpretation alone. A series of interpretive studies are planned to identify the optimal interface between “black box” outputs and radiologist interpreters, which remains an understudied topic. Our new end-user tool aims to help tip the balance of routine screening towards greater benefits than harm.